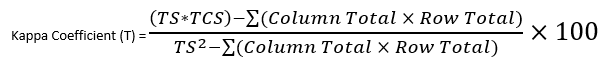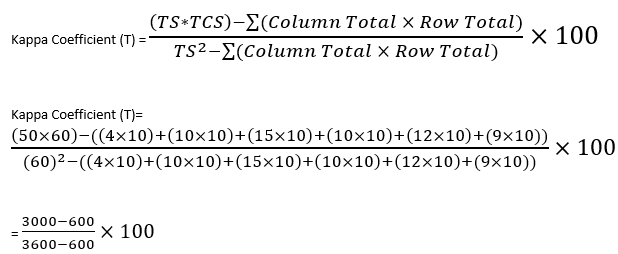Random Forest Supervised Classification Using Sentinel-2 Data
Contents
Introduction to Multi-spectral Imaging
Multispectral imaging (MSI) captures image data within specific wavelengths ranges across electromagnetic spectrum. MSI detects different images through instruments that are sensitive to different wavelengths of light thus allowing for distinction in land-type. MSI is a highly informative form of imaging technique as it can move beyond visible light range and can detect and extract data that the human eye fails to capture. Sentinel 2 is an Earth Observation mission from the Copernicus programme that acquires high resolutions multispectral imagery by conducting frequent visits over a given area. Sentinel-2 is a polar-orbiting Earth Observation Mission from the Copernicus programme that conducts multispectral high-resolution imaging for land monitoring to provide, for example, imagery of vegetation, soil and water cover, inland waterways, and coastal areas. Sentinel-2 can also deliver information for emergency services. Sentinel-2A was launched on 23 June 2015 and Sentinel-2B followed on 7 March 2017. The Sentinel-2 has 13 bands of multispectral data in the visible, near infrared and short-wave infrared part of the spectrum. This tutorial applies images captured by the Sentinel 2A satellite, which provides similar functions to the ones mentioned above.
Supervised and Random Forest Classification
Supervised Classification is a technique used for extracting information from image data. The process includes classification of pixels of an image into different classes based on features of the pixels. Supervised classification can be conducted in 2 main stages. The first stage is called Training Stage and the latter is called Classification Stage. In the training stage of the process, a set of vectors called training sample is established by the user through which the supervised classification is conducted. The number of classifiers is depended upon the number of inputs by the user. For example, if the user identifies 5 different land cover classes in its training data, the supervised classification conducted will output an image of the scene using the data and will have 5 different class distinctions. This tutorial uses a specific type of supervised classification technique called Random Forest.
Random Forest Classification is a supervised form of classification and regression. Random forest or random decision forests are a form of Model Ensembling techniques. Model Ensembling attempts to aggregate large number of test models to improve accuracy in classification and regression. Random Forest works on the principle of Model Ensembling and helps provide accurate low-cost classified images using a baseline of training data. Higher the training data for classification, higher the accuracy in the end product.
Software and Data Acquisition
This section provides information on the software's and data to be used to conduct the processes listed in the tutorial.
Sentinel Application Platform (SNAP)
The Sentinel Application Platform - or SNAP - in short is a collection of executable tools and Application Programming Interfaces (APIs) which have been developed to facilitate the utilisation, viewing and processing of a variety of remotely sensed data. The functionality of SNAP is accessed through the Sentinel Toolbox. The purpose of the Sentinel Toolbox is not to duplicate existing commercial packages, but to complement them with functions dedicated to the handling of data products of earth observing satellites.
The following tutorial uses SNAP Version 8.0. The software is open source and can be downloaded through the following link: http://step.esa.int/main/download/snap-download/. Specific toolboxes for data processing of data from different Sentinel Satellites can also be downloaded through the link.
This tutorial uses a Windows 64-bit operating system with Windows 11 and 16GB of RAM. The software can also run on a Mac OS X, Linux and 64-bit and 32-Bit Windows. The software recommends having at least 4GB of memory. To run the 3D World Wind View, it is recommended to have a 3D graphics card with updated drivers. However, this tutorial will not be using 3D World Wing View.
Microsoft Excel / OpenOffice Calc
The tutorial also uses Microsoft Excel Software to Conduct Kappa Coefficient Analysis. The software can be downloaded from the link: https://www.microsoft.com/en-ca/microsoft-365/p/excel/cfq7ttc0k7dx/?activetab=pivot%3aoverviewtab
However, the software is only available through purchase therefore an alternative software to conduct similar processes is Microsoft OpenOffice Calc. This software is free to use and can be downloaded from the link: https://openoffice.en.softonic.com/download
Tutorial and SNAP Data
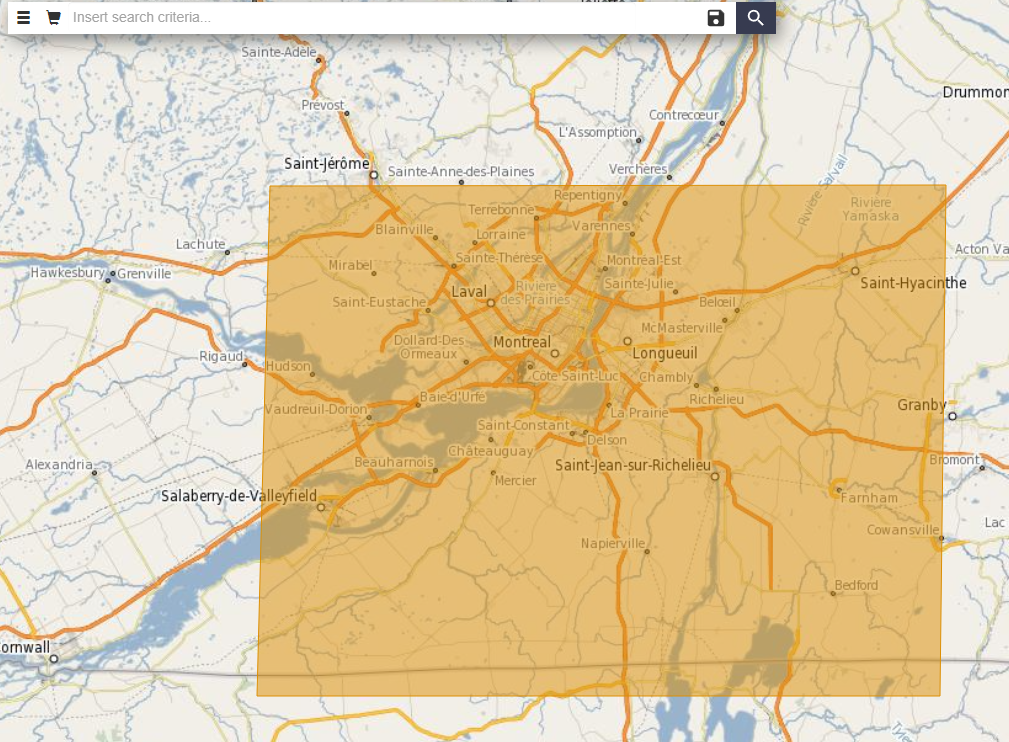
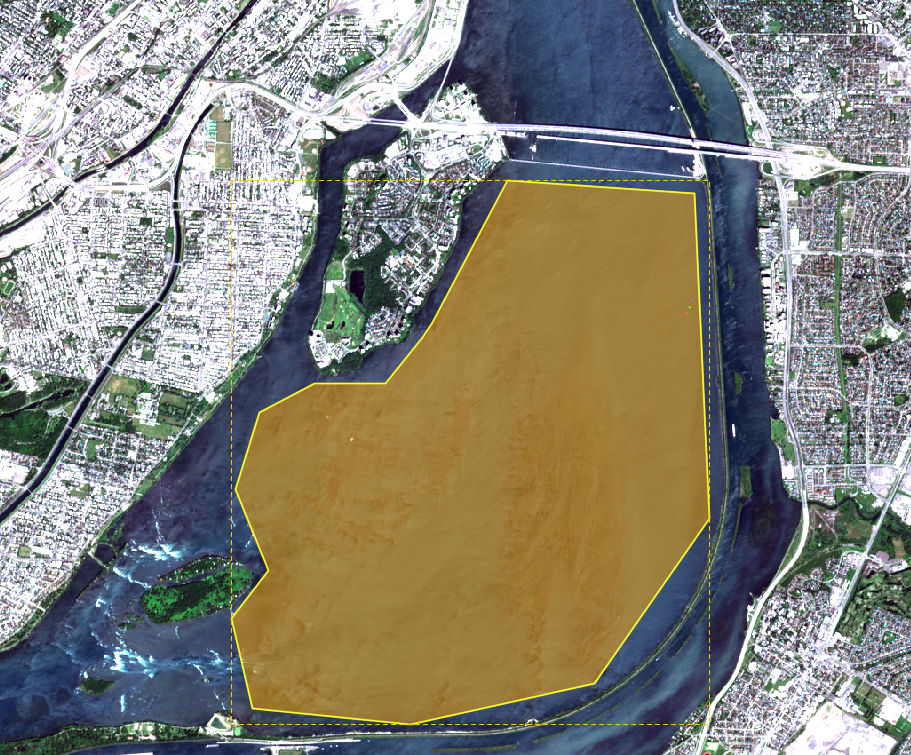
To acquire the data used in the tutorial, you will have to create a free account with Scihub, also known as, Copernicus Open Access Hub. The free to use and open access hub allows access to Sentinel 1, 2, 3, 5P data. Go to the link: https://scihub.copernicus.eu/dhus/#/home and to the top right corner, you will find a symbol of account login. Below that you will find a link to “Sign Up”. Click that and fill out the information. Once your account is set up, you can begin by entering the same link and typing in the search bar, the type of data you wish to download. A different approach through which data can be acquired is by creating a polygon space over the area you desire to use for the tutorial. Hover over the map and zoom In or Out depending on the scale of the area you wish to use. Once you have the extent you wish to use, CLICK the Right Pointer on your Mouse. A line will appear, extend the line to your liking, and CLICK the Right Pointer again and now you will have a 2-side completed polygon. Repeat the steps until you have a complete polygon, once that is achieved, DOUBLE CLICK the right button on your Mouse, this will concretize your area of interest. Refer to Figure 1 Below for guidance on how the polygon will look.
Figure 1: Screenshot of the polygon space created over AOI.
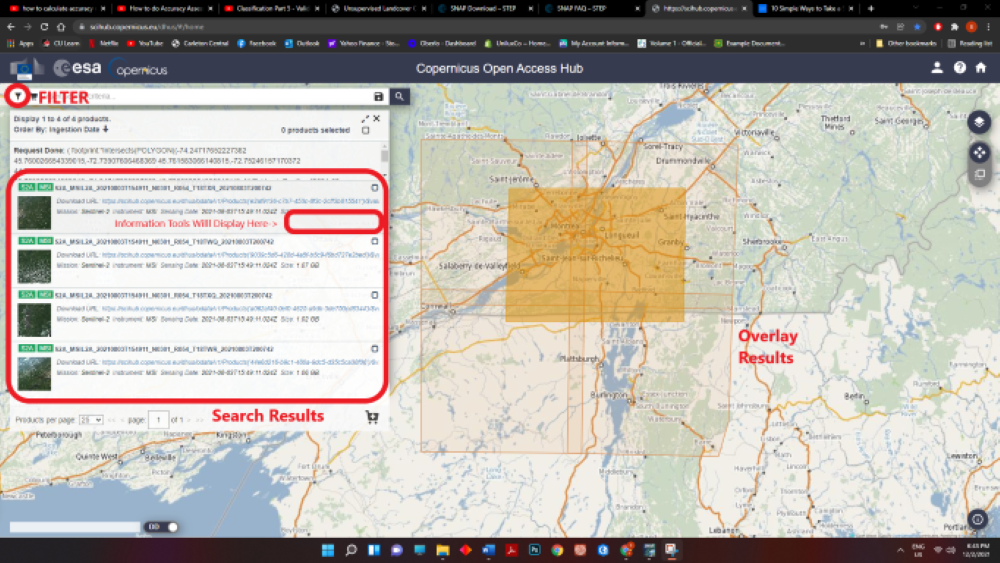
Next, Click the SEARCH button near the search bar. You will now have different polygon overlays of imagery data acquired from different Sentinel Satellites.
Figure 2: Screenshot of the window with search results and filter button navigation.
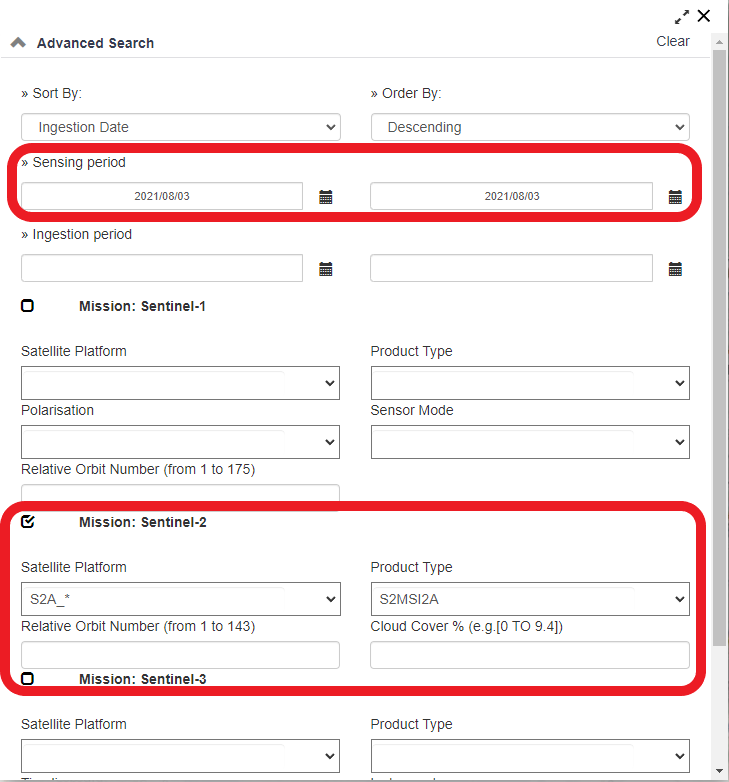
To further narrow you search to specific dates and Satellite Types, Click the FITLER tab and change the settings according to your wishes. The window will look like the Figure 3 below:
Figure 3: Screenshot of the filter window tab.
The data used in this tutorial cab be attained using the following parameters in the filter tab:
Location: Montreal
Sensing Period: 08/03/2021-08/03/2021
Satellite Type: Sentinel 2A
Product Type: MSI2A
Dataset Name: S2A_MSIL2A_20210803T154911_N0301_R054_T18TXR_20210803T200742
This can also be found in the Figure 3. Once the filters have been applied. The best overlaying polygon (one that covers all/or most of the area of interest) can then be downloaded through hovering over the result and clicking the DOWNLOAD ARROW SYMBOL on the BOTTOM RIGHT of the Search Result.
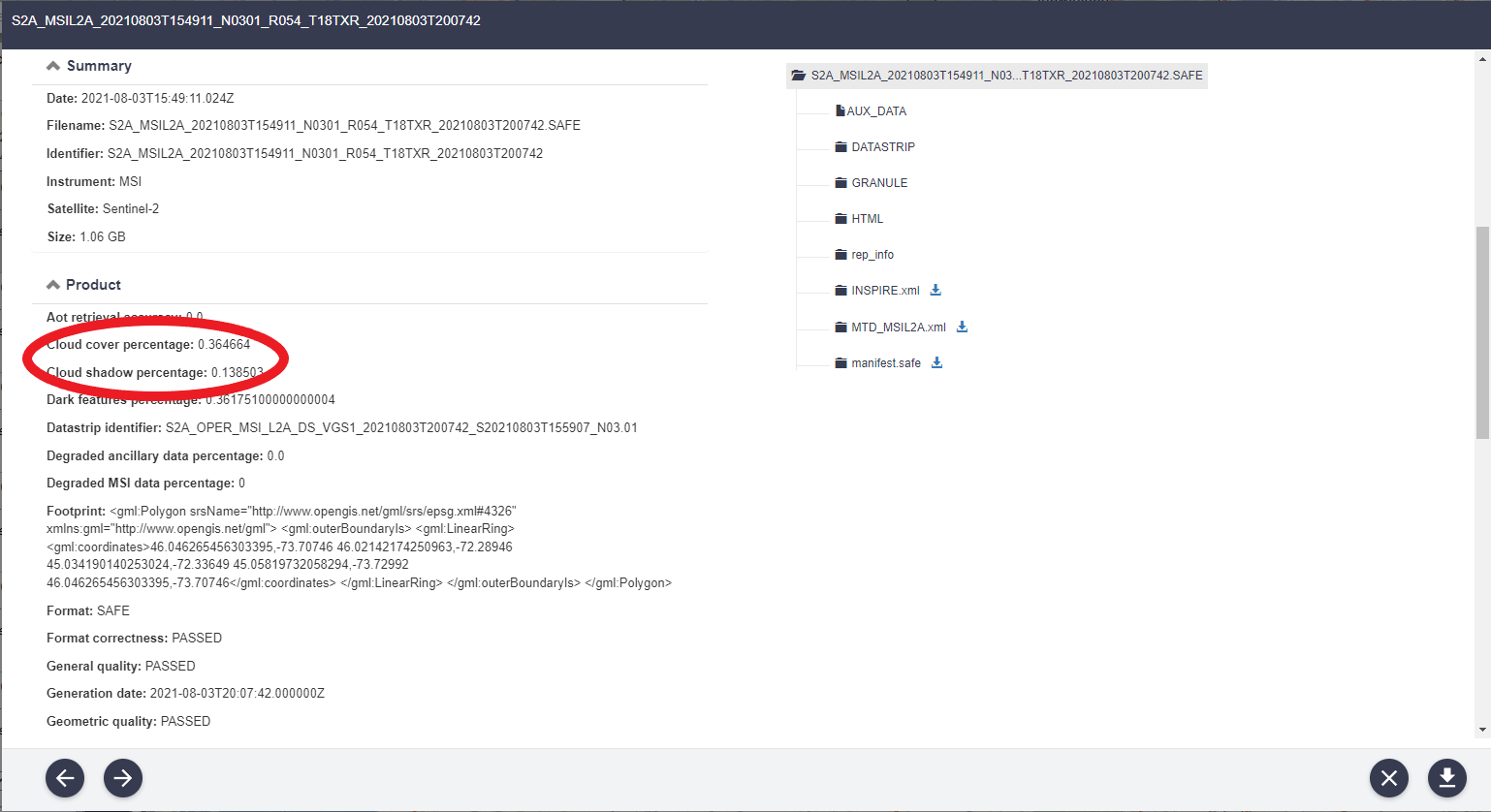
It is important to attain data that has the least amount of cloud cover. This information can be found by clicking the EYE SYMBOL beside the sensing output search. Under PRODUCT INFORMATION, you will find the CLOUD COVER PERCENTAGE. This value should be near 0 as this allows for a more accurate SUPERVISED CLASSIFICATION. An example of this information can be found in the Figure 4 below:
Figure 4: Screenshot of the product details tab, displaying the cloud cover percentage of the acquired image.
Preprocessing Data and Workspace
How to Toggle Viewing Data in SNAP
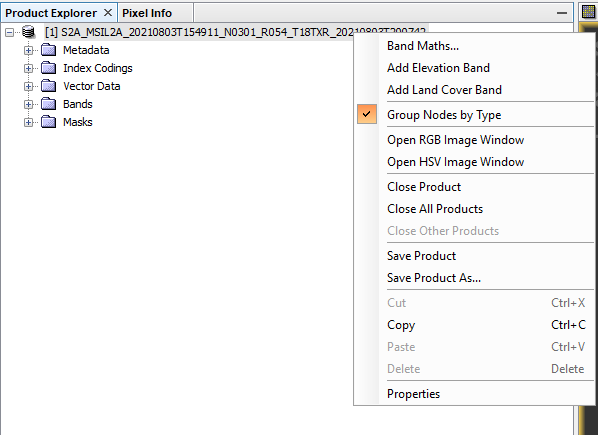
To begin conducting the classification you first need to establish a workspace to safely store and access your data during and after you classification analysis. Begin by opening SNAP and importing the folder with the data you have acquired. This will prompt the folder of data and the corresponding bands, vector data, metadata, masks, and index coding. For this tutorial we will only be working with the bands and vector folders.
To open and look through your data:
To view Natural Image: Right click the data folder --> click OPEN RGB IMAGE WINDOW --> from the scroll down tab, choose MSI NATURAL COLORS --> Click OK.
To view the infrared image: open the RGB IMAGE WINDOW --> from the scroll down tab chose FALSE COLOR INFRARED --> Click OK. This can be referred to visually through the Figure 5 below:
Figure 5: Displays how to access the RGB Image Window in SNAP.
To toggle and move around your screen when zoomed into your scenery, Use the tool from the Quick Access Toolbar that looks like a ‘Hand’. It should look something like this:
Figure 6: Displays the tool for viewing and moving across the map.
Both of these processes will pop up the natural and false color infrared images respectively.
Preprocessing
The tutorial utilizes Sentinel 2 data, this data is of Level-2A processing and includes a scene classification and an atmospheric correction applied to Top-Of-Atmosphere (TOA) Level-1C orthoimage products. Level-2A main output is an orthoimage Bottom-Of-Atmosphere (BOA) corrected reflectance product. Therefore, extensive pre-processing steps are not required for the data at hand, only a resampling and reprojection is required to conduct the assessment (Steps Below). Although if you wish to use other levels of data, the following link of blog from the Freie University, Berlin, can help you understand the different pre-processing steps: https://blogs.fu-berlin.de/reseda/sentinel-2-preprocessing/.
S2 Resampling:
Sentinel 2 data has different bands under various resolutions. To conduct processes in latter parts, the data needs to be at a similar resolution. Therefore, resampling of the different bands is required.
To conduct the resampling follow the steps below:
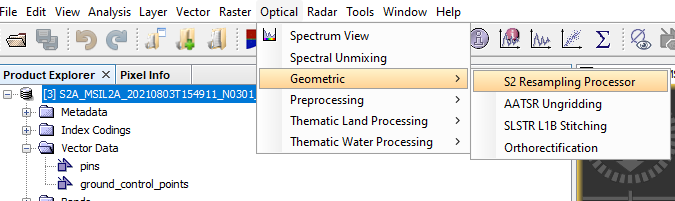
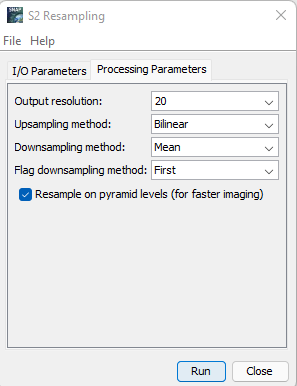
Open Optical from the navigation bar at the top --> Open Processing --> Open S2 Resampling --> Given an Appropriate name to your S2 Resample Result --> Select the Main Imagery as the Input --> Open the Parameters Section from the Menu --> Set the Resolution to 20m --> Run.
The steps can be seen in the figure below:
Figure 7: Displays how to open S2 Resampling Processor.
Figure 8: Displays the parameters to enter in the S2 Resampling Processor.
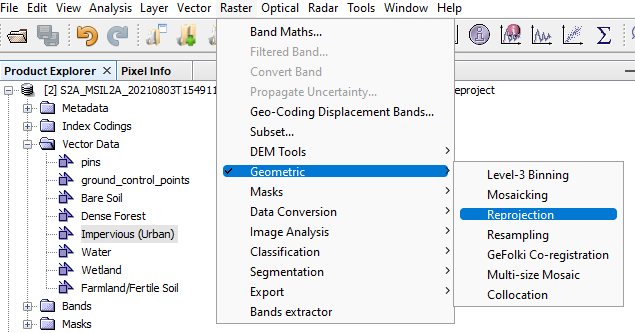
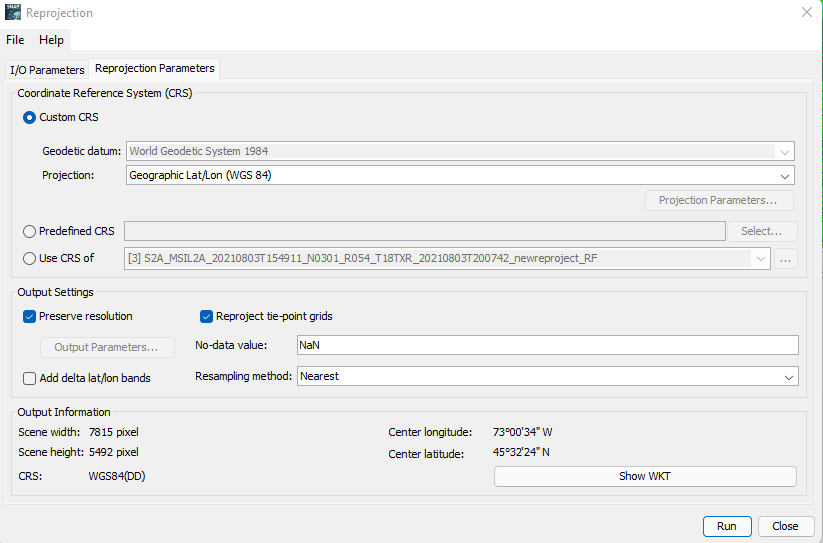
Reprojection: Although the data itself does not need preprocessing, SNAP due to its processing issues in classification only allows data with WGS 84 projection to be applied. To reproject your data: Raster --> Geometric --> Reprojection --> Set your Input as the S2 Resampled Layer Acquired from the previous step and give an appropriate name to your output --> Open Parameters and set the projection to Geographic Lat/Lon (WGS 84) --> Run
The steps and the parameters are shown in the figures below:
Figure 9: How to open Reprojection tool window.
Figure 10: Displays the parameters to enter in the Reprojection tool.
Developing Training Data
Creating Vector Containers
To conduct our supervised classification, we will need vector training data. This data will help establish the different classes for our supervised classification. Therefore, we need vector containers to develop the training data.
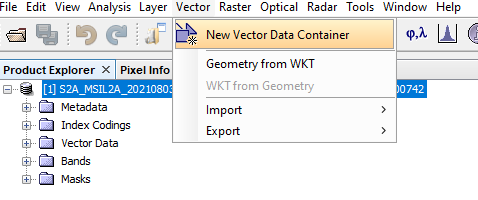
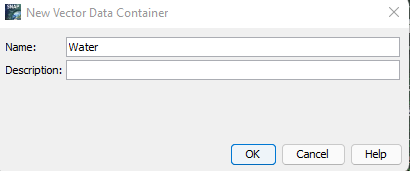
To build vector containers: From the Navigation bar, Select Vector --> New Vector Container. Name this according to the first class of data you would like to establish. This tutorial first develops Water Data. Therefore, in the Name of the NEW Vector Container, you write Water. The steps can be referenced from figures 11 and 12 below:
Figure 11: Displays how to create new vector container
Figure 12: Displays how the vector data will be named
Creating Vector Features
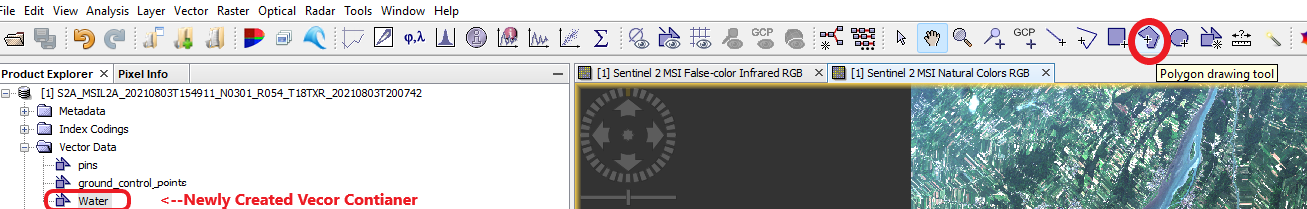
Begin by hovering over and zooming into a section of water features in the spectral image. Go to Vector Data on your working Folder --> Choose the newly created Water Vector Container Once you have clicked the container, near the top right of the page you will see a function called CREATE POLYGON FEATURES. An example of this function can be seen in figure 13 below:
Figure 13: Displays navigation to Create Polygon Features tool and shows the newly created vector container.
CLICK the polygon features tab, Hover to the area with water and add 4 or more points to make a polygon, once a polygon is created DOUBLE CLICK the polygon to concretize it. Refer to figure 14 below:
Figure 14: Displays how vector training features are created.
Now you have created a polygon feature that will act as the training data to discern water in the supervised classification. Create as many water features as you would like. For this tutorial, the user should aim to add at least 10 feature polygons for each vector container/feature they deem to add in the classification.
Note: It is important to note that as many features/training polygons you will add in the vector containers, the more accurate will the outcome of the supervised classification will be. However you have to be careful of the pixels you incorporate in the vector container classes as diluted polygon features can cause your classification to be confused.
There can be many different shades of the same feature, for example the spectral image at hand has many different shades of water across. Therefore, it is important to add different shades of the same feature. Refer the figure 15 below for a better understanding of how to add different feature polygons.
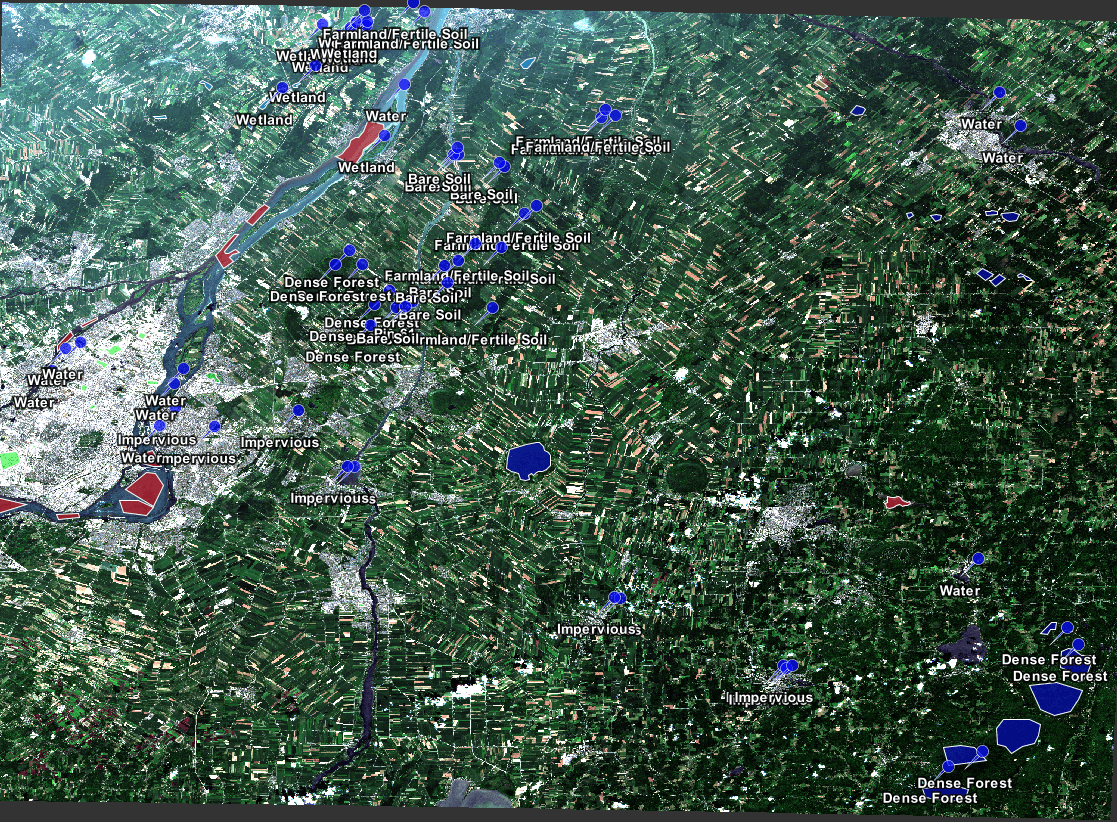
Figure 15: Displays how the vector training data is scattered across the imagery.
To understand the spectral image better and to help you choose the different vector container classes, you can make use of the scatterplot. Scatterplot of a vector container helps you understand the wavelength at which the data you have added lies.
To access scatterplots of your own, Go to ANALYSIS in Navigation toolbar --> Choose SCATTERPLOT --> For the ROI Mask, Choose Water --> For X-axis choose RED (B4) and for Y-Axis Choose NIR 1 (B8). Below is an example of scatterplot created from Water Vector Features.
Figure 16: Displays how to access the scatterplot
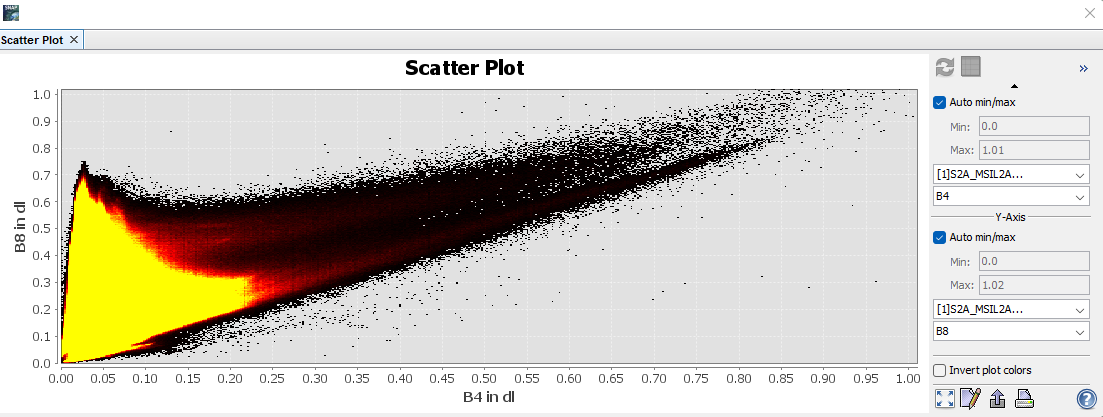
Figure 17: Displays the parameters and resulting scatterplot for water vector training features.
It can be seen from Figure 17 that most of the training polygons lie in the bottom left of the scatter plot i.e., where they should be as they are the lowest wavelength bands.
Sometimes features can be hard to discern. To mitigate this issue, you can also toggle the viewing image you have in front of you. If you are using a Natural True Color Image, change over to the NEAR INFRARED image. This image will help discern small changes in the land cover type in cases of bare soil, vegetation, and dense forests. Follow the steps stated in the data and SNAP toggling section. The resulting NEAR-INFRARED image should look something like this:
Figure 18: Displays the near-infrared image of the chosen imagery.
It can be seen from the Figure 18 above that distinction between different classes has now become easier as the near-infrared image allows for small changes to be seen that the naked eye might miss from the natural color imagery.
Now, use all of the information provided above and add as many feature classes as you like. The following tutorial creates a total of 6 different feature classes for vector containers with 10 vector samples for each. The classes created are: Water, Wetland, Bare Soil, Urban Areas, Farmland/Fertile Soil and Dense Forest.
Random Forest Classification
Now that you have all the required vector classes inserted with feature polygons, it is time to conduct the random forest classification.
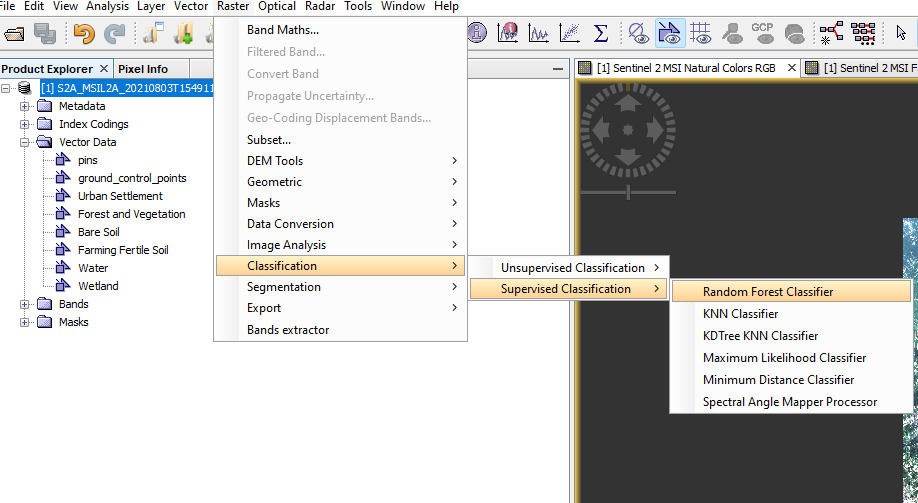
Head over to ‘Raster’ --> Classification --> Supervised Classification --> Random Forest Classifier. It can be referred from the figure below:
Figure 19: Displays how to access Random Forest Supervised Classification in SNAP.
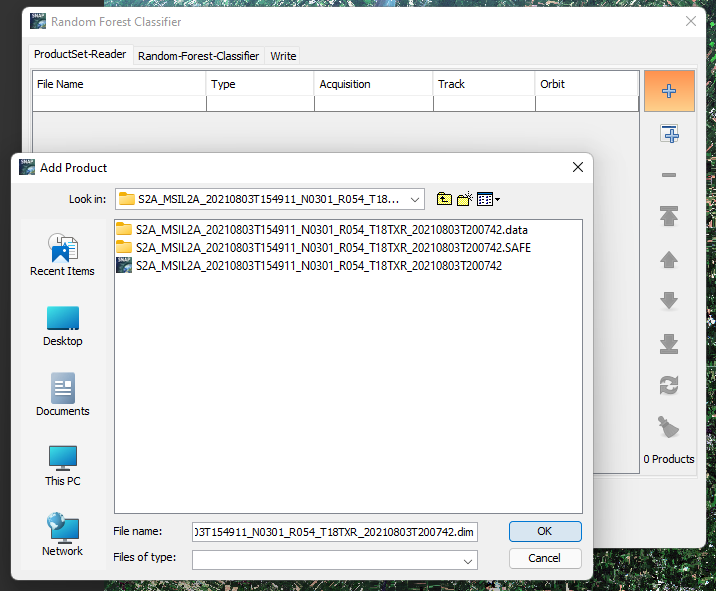
To add your data click the add current map button on the side menu, You can refer to it in the figure below:
Figure 20: Displays how to add data in Random Forest Classifier.
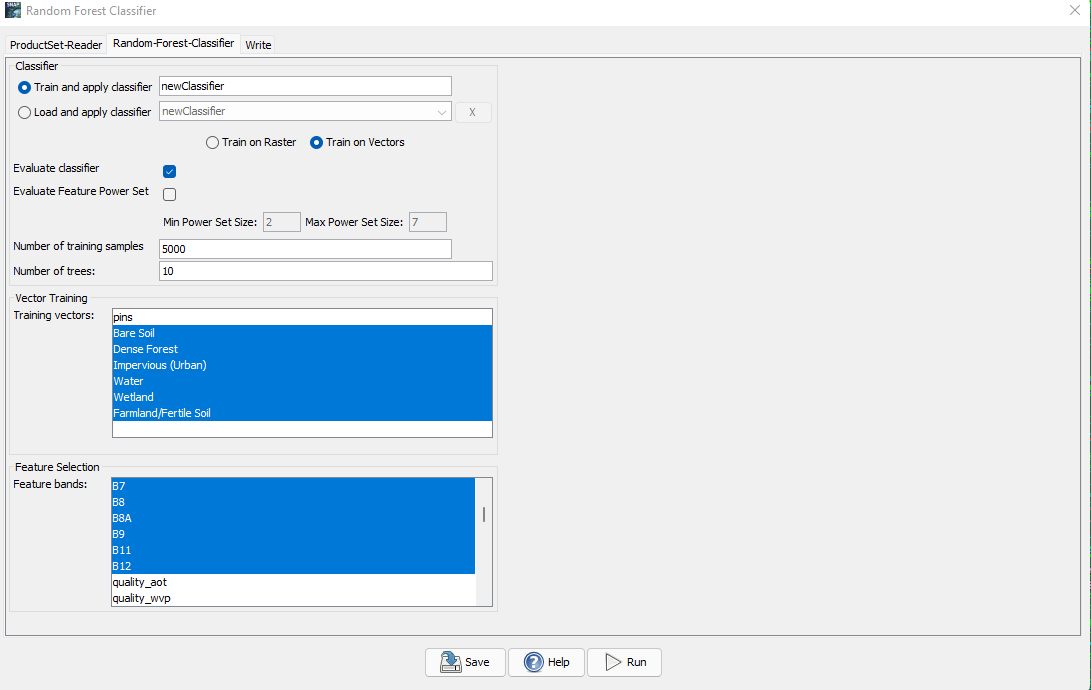
For the classification we will be using only Bands 1-12. Using additional bands can help increase accuracy as it will provide the classification more sceneries and data to compare. However, this process can take time and since the tutorial is only an introductory tutorial to random forest classification, we will only be using bands 1-12. Now follow the same parameters installed in the following figure and run:
Figure 21: Displays the parameters to choose for random forest classifier.
To view your resulting classification, go to the output project and open the ‘BANDS’ folder, here now, instead of different frequency bands, you should have two different outputs: “Labeled Classes” and “Confidence”. Go ahead and DOUBLE CLICK the Labeled Classes Output. A new window should open, and the output of the classification should look something like this:
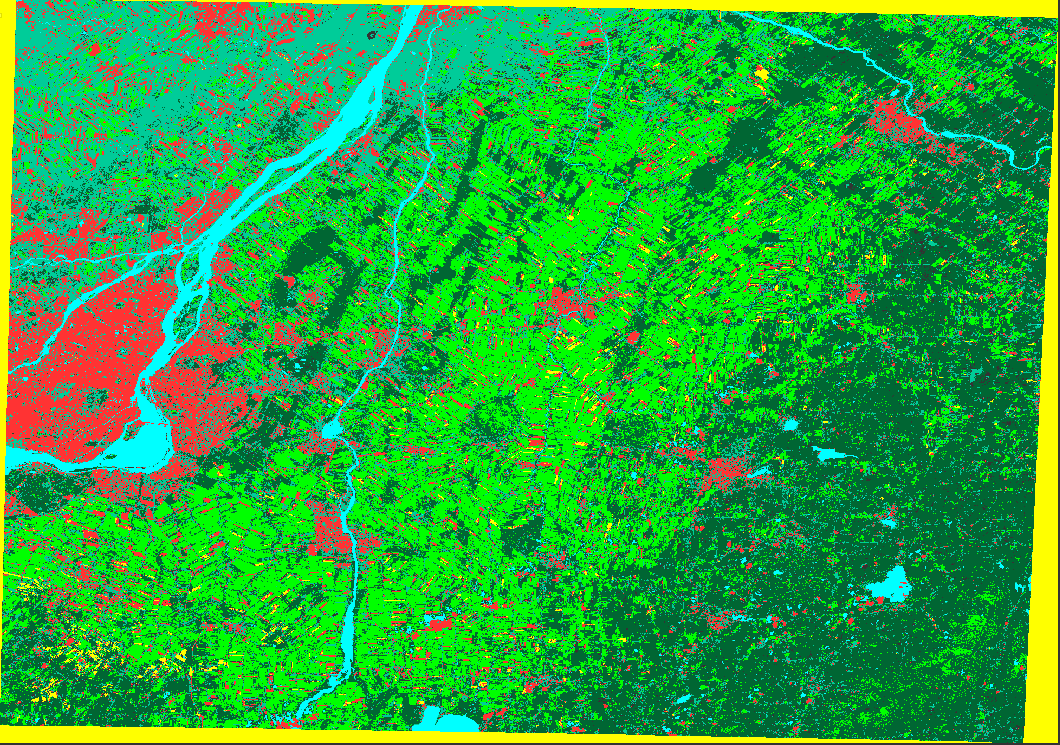
Figure 22: Displays the results of the classification.
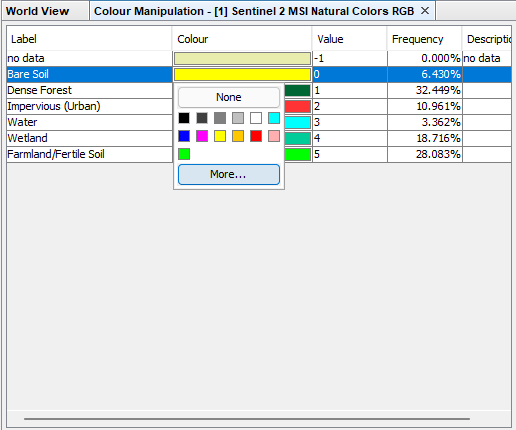
To make the classes clearer to yourself you can also change the colors of the classes in front of you. Simply, Open Color Manipulator Toolbar, the symbol should be in the Quick Access Toolbar, and it should look something like this:
Figure 23: Displays the color manipulator tab.
You can change the colors of each output class according to your liking by DOUBLE CLICKING the color bar and choosing from the options available.
Now you have a more distinct and readable classification output.
Validation
Preparing for Validation
To conduct our validation of the classification results, we will be using the manual version of the kappa coefficient method.
To conduct this process, we first need to place different pins of data on our ORIGINAL MAP (The one that is resampled and reprojected.) to coagulate the different points to the results of the classification. We will be placing over 10 different pins for each class. Normally this number should be higher to have a robust validation method but for the sake of this tutorial we will only be applying 10 pins for each. To place different pins, from the Quick Access Toolbar, Click the “Pin Placing Tool”, situated beside the search symbol.
Now use the “Hand” tool to move around the map and begin with the feature that is labeled as 0 in your classification. This can be found in the Color Manipulator Toolbar. Place 10 pins each. Remember to switch between bands when starting to place pins for a new class. If this is not done the new pin you input will be accidently added in the land class you were using earlier. Once that is done, you need to label each of these pins according to their feature for further examination.
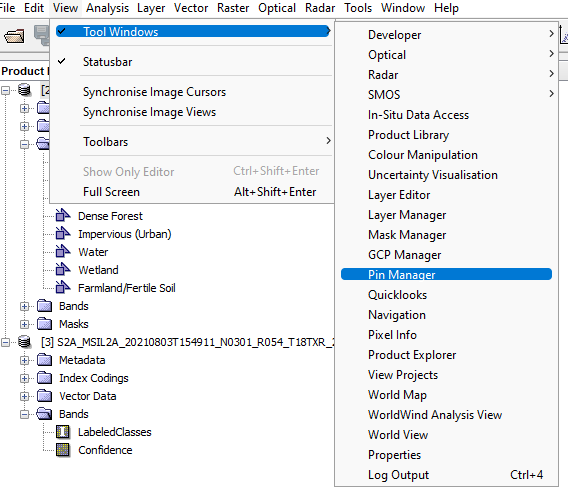
To do this go to VIEW --> TOOL WINDOWS --> PIN MANAGER. Follow the process shown in the figure below:
Figure 24: How to access the pin manager toolbar.
Here you can change the name of the pin according to your placing in the ‘Labeled’ Column by DOUBLE CLICKING it. Label each 10 pins according to their feature class i.e.: water, bare soil, vegetation etc. Make sure to place the pins apart and provide some distinction across the map as this will provide your data to be more thoroughly evaluated. The map should look something like this:
Figure 25: Shows the diversity of pin placement.
You should have a total of 60 pins if you have 6 classes and more if you have chosen to have more feature classes. Now we need to compare these pins to the classified data. To do this you need to transfer the pins from the ORIGINAL MAP to the CLASSIFIED MAP.
Go to the PIN MANAGER menu, on the right-hand side, you will see a symbol similar to this:
Figure 26: Display the tool for transferring data.
This button will transfer the pins from your ORIGINAL MAP to CLASSIFIED MAP for comparison. Select all the pins by pressing Ctrl + A and then click the symbol and a window will open asking which project to transfer the pins to. Choose you RANDOM FOREST CLASSIFIED map and click OK.
Now open your CLASSIFIED MAP and confirm that all the pins have been transferred by opening the PIN MANAGER for the project again.
As mentioned, we need to compare the two results. One was chosen by you, by looking at the original map and eyeballing your data for comparison through a remotely sensed image and the other was chosen through training data. SNAP has an important and useful feature that allows the comparison of the results.
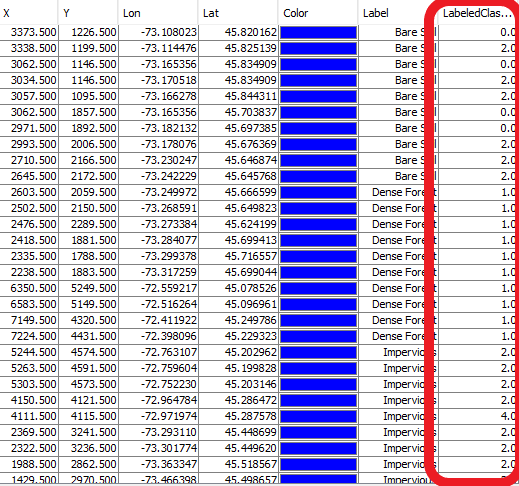
The PINS that have been placed will help conduct this. In the PINS MANAGER TOOLS for your CLASSIFIED MAP, you will see a tool called “Filter Pixel Data” next to the tool you used earlier for transferring data. This will use the assigned classification values from your result and apply them into a new column for the pins you have chosen for training into the PINS table that you transferred. The pixel assigned values table should look something like this:
Figure 27: Displays the output table for filter pixel data in pins manager.
Contingency Table and Kappa Coefficient Analysis
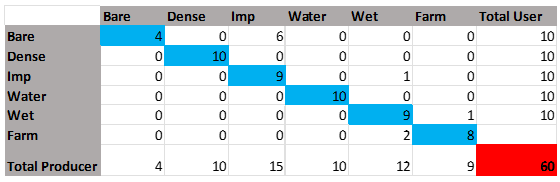
Now in order to test our model’s accuracy, we can evaluate the errors and calculate the overall accuracy and kappa coefficient. To do this we can use Microsoft Excel or OpenOffice and numerate the data we gathered. Below is a contingency table that will help us conduct the calculations for overall accuracy and the kappa coefficient:
Figure 28: Displays the Kappa Coefficient Analysis Table.
Place all your features according to the table displayed above and create more columns and rows if needed. The “USER VALUE” in this case is the total number of pins we have placed for each feature, and this will stay constant across the column. Now we can begin using the PINS MANAGER TABLE we have in SNAP. Sort the table by clicking the “LABEL” column as this will help keep similar data together.
The “LABELED CLASSES” column in the CLASSIFIED MAP table presents the pixel value i.e., classified location value from the classification results. In the first part of the table i.e., Bare soil, calculate the number of RIGHT CLASSIFICATIONS. In the case of the table used in the tutorial, Bare Soil is indicated by the ‘Value’ ‘0’. Refer to the figure 23 above.
You will see that out of the 10 validation pins placed for bare soil, only 4 have been correctly identified. This value will be inserted in the EXCEL TABLE’S top compartment where both column and row have a similar heading i.e., Bare Soil (Figure 28). Identify what other pins have been incorrectly identified as, which in this case, 6 pins have been identified as impervious land. Insert that value into the impervious column crossing the bare soil row in the table. Continue this process and fill out the rest of the table. Remember that all the correctly identified PIN numbers need to flow diagonally and all other incorrectly identified PINS should be placed according to their identification.
The TOTAL PRODUCER value now needs to be calculated and it can be done by adding all the values from each column. When all the diagonal values are summed, the total is 50 which when inserted into a percentage equation i.e. (50/60)*100 provides us an accuracy of 83%. However this only displays the correctly identified accuracy, to measure the entire model i.e. CLASSIFICATION RESULT, we insert the data developed in the table above into a kappa coefficient equation.
The Equation can be found below:
Figure 29: Displays the kappa coefficient equation.
In this equation TS is the total number of samples i.e., 60 and TCS is the Total number of Correctly Identified Samples. Follow the formula and insert the values you have received from your table and calculate like the one conducted below:
Figure 30: Displays the calculations for the Kappa Coefficient Analysis.
Therefore the overall accuracy of the Random Forest Supervised Classification is found to be 80%.
Conclusion
By completing this tutorial, you will be able to execute random forest supervised classification and validation using only the SNAP platform. However, there are many other integrations that can be applied to substitute some of the processes conducted in the tutorial. The overall accuracy of the resulting classification was found to be 80%, which can be increased by adding more training data and data that does not coincide with the classes chosen for the classification. SNAP is a really powerful earth observation processing and analysis tool. It has many capabilities, one of which has been explored by this tutorial. The knowledge acquired through this tutorial will help you execute quick and easy image processing tasks.
References:
- Delgato, Montessarrat, and Joaquin Sendra. “Validation of GIS-Performed Analysis - Researchgate.net.” Validation of GIS-performed analysis. Research Gate, January 2009. https://www.researchgate.net/publication/287541349_Validation_of_GIS-performed_analysis.
- Pulsifier, Peter, and Scott Mitchell. “Uncertanity, Error and Accuracy in GIS: Overview.” Lecture, September 22, 2021.
- Siegel, Jon, and Richard Mark Soley. “Open Source and Open Standards: Working Together for Effective Software Development and Distribution.” Open Source Business Resource, November 1, 2008. https://timreview.ca/article/207#:~:text=Open%20source%20refers%20to%20software,of%20software%20or%20software%20interfaces.
- Sekulić, Aleksandar, Milan Kilibarda, Gerard B.M. Heuvelink, Mladen Nikolić, and Branislav Bajat. 2020. "Random Forest Spatial Interpolation" Remote Sensing 12, no. 10: 1687. https://doi.org/10.3390/rs12101687
- “SNAP.” STEP. Accessed December 23, 2021. https://step.esa.int/main/toolboxes/snap/